Beyond the Bench
Cancer Research Highlights
The University of Kansas Cancer Center aims to do nothing less than fundamentally transform the cancer experience – to give all people in our region greater access to breakthrough ideas and novel treatments.
From deepening our understanding of cancer's biology to expanding cancer prevention efforts to ensuring patients' care continues after treatment, we are tackling cancer on multiple fronts. Beyond the Bench, an annual publication, highlights some of the latest, most innovative research efforts happening at the cancer center.
Read Beyond the Bench
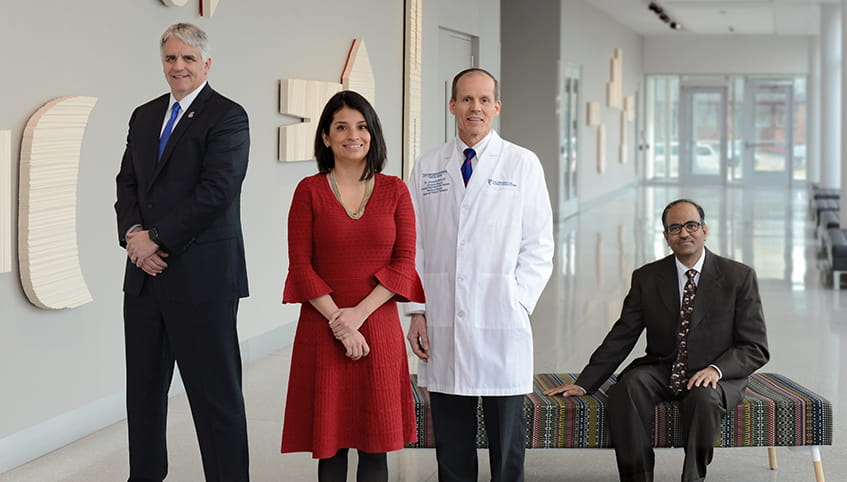


NCI Designation
Our cancer center is 1 of fewer than 60 National Cancer Institute-designated comprehensive cancer centers nationwide. 
Clinical Trials
We are a pioneer in new clinical trial research that leads to innovative treatments. 
Still the Best
Our hospital continues to rank as the best in Kansas City and in Kansas
according to U.S. News & World Report.